User login
Product gets orphan designation for AML
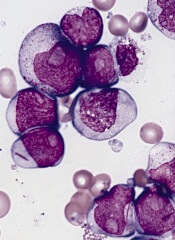
The US Food and Drug Administration (FDA) has granted orphan designation for eryaspase to treat acute myeloid leukemia (AML).
Eryaspase, also known as ERY-ASP or GRASPA, is L-asparaginase encapsulated in red blood cells.
These donor-derived, enzyme-loaded red blood cells function as bioreactors to eliminate circulating asparagine and “starve”
leukemic cells, thereby inducing their death.
Research has suggested this delivery system provides improved pharmacodynamics. It protects L-aspariginase from circulating proteolytic enzymes and prevents early liver or renal clearance.
The system also appears to reduce the risk of adverse events.
Eryaspase is currently under investigation in a phase 3 trial for acute lymphoblastic leukemia (ALL) and a phase 2b trial for AML in Europe. A phase 1 study in adult ALL is being launched in the US.
Eryaspase now has orphan designation for ALL, AML and pancreatic cancer, both in Europe and the US.
In the US, orphan designation is generally granted for drugs or biologics intended to treat disorders of high unmet medical need that affect fewer than 200,000 people.
This designation conveys special incentives to the product’s sponsor, including 7 years of US market exclusivity for the drug or biologic upon FDA approval, a prescription drug user fee waiver, and certain tax credits. ![]()

The US Food and Drug Administration (FDA) has granted orphan designation for eryaspase to treat acute myeloid leukemia (AML).
Eryaspase, also known as ERY-ASP or GRASPA, is L-asparaginase encapsulated in red blood cells.
These donor-derived, enzyme-loaded red blood cells function as bioreactors to eliminate circulating asparagine and “starve”
leukemic cells, thereby inducing their death.
Research has suggested this delivery system provides improved pharmacodynamics. It protects L-aspariginase from circulating proteolytic enzymes and prevents early liver or renal clearance.
The system also appears to reduce the risk of adverse events.
Eryaspase is currently under investigation in a phase 3 trial for acute lymphoblastic leukemia (ALL) and a phase 2b trial for AML in Europe. A phase 1 study in adult ALL is being launched in the US.
Eryaspase now has orphan designation for ALL, AML and pancreatic cancer, both in Europe and the US.
In the US, orphan designation is generally granted for drugs or biologics intended to treat disorders of high unmet medical need that affect fewer than 200,000 people.
This designation conveys special incentives to the product’s sponsor, including 7 years of US market exclusivity for the drug or biologic upon FDA approval, a prescription drug user fee waiver, and certain tax credits. ![]()

The US Food and Drug Administration (FDA) has granted orphan designation for eryaspase to treat acute myeloid leukemia (AML).
Eryaspase, also known as ERY-ASP or GRASPA, is L-asparaginase encapsulated in red blood cells.
These donor-derived, enzyme-loaded red blood cells function as bioreactors to eliminate circulating asparagine and “starve”
leukemic cells, thereby inducing their death.
Research has suggested this delivery system provides improved pharmacodynamics. It protects L-aspariginase from circulating proteolytic enzymes and prevents early liver or renal clearance.
The system also appears to reduce the risk of adverse events.
Eryaspase is currently under investigation in a phase 3 trial for acute lymphoblastic leukemia (ALL) and a phase 2b trial for AML in Europe. A phase 1 study in adult ALL is being launched in the US.
Eryaspase now has orphan designation for ALL, AML and pancreatic cancer, both in Europe and the US.
In the US, orphan designation is generally granted for drugs or biologics intended to treat disorders of high unmet medical need that affect fewer than 200,000 people.
This designation conveys special incentives to the product’s sponsor, including 7 years of US market exclusivity for the drug or biologic upon FDA approval, a prescription drug user fee waiver, and certain tax credits. ![]()
Testing reveals abnormalities in CN-AML/MDS

Credit: NIGMS
NASHVILLE—New research suggests we may need to use more sensitive methods to analyze patients with cytogenetically normal acute myeloid leukemia or myelodysplastic syndrome (CN-AML/MDS).
Using “highly sensitive” microarray technology, researchers found a distinct pattern of genetic abnormalities in 22 patients diagnosed with CN-AML/MDS.
The team identified 3 overlapping regions of homozygosity in 3 genes, 2 of which are known to be involved in carcinogenesis.
This suggests that using karyotyping or FISH, or simply looking for known mutations, is not sufficient for evaluating patients with CN-AML/MDS, according to Ravindra Kolhe, MD, PhD, of the Medical College of Georgia at Georgia Regents University.
“The technology we currently use can’t identify specifically what’s wrong,” Dr Kolhe said. “We have to use more sensitive tests to give patients the proper answer.”
Dr Kolhe presented this finding, and the research to support it, at the American College of Medical Genetics and Genomics Annual Clinical Genetics Meeting.
He and his colleagues analyzed 22 patients. Seventeen had AML, and 5 had MDS, including 1 with refractory anemia with excess blasts-2. All patients had normal karyotype and FISH and had greater than 20% blasts in the bone marrow.
The researchers analyzed samples from these patients using a high-resolution, single-nucleotide polymorphism (SNP) microarray called CytoScanHD.
According to the company that markets this technology (Affymetrix, Inc.), the assay includes 750,000 SNPs with over 99% accuracy to detect accurate breakpoint estimation, loss of heterozygosity determination, regions identical-by-descent, maternal contamination, and low-level mosaicism.
For Dr Kolhe and his colleagues, the assay revealed small, previously undetectable changes in patients thought to be cytogenetically normal.
Specifically, the researchers identified 3 overlapping regions of homozygosity in all 22 cases—chromosome 1p34.3, chromosome 1p32.3, and chromosome 16q22.1 in the SFPQ, EPS15, and CTCF genes, respectively.
SFPQ and CTCF are already known to be involved in carcinogenesis, and Dr Kolhe and his colleagues are now investigating the role of EPS15 in leukemogenesis.
The researchers also identified additional abnormalities and are investigating these as well. They are sequencing the genes to identify homozygous or compound heterozygous mutations, performing expression studies to confirm that these mutations are leukemic, and conducting experiments in knockout mice to demonstrate that these genes produce the same leukemia phenotype.
The materials and reagents for this study were provided by Affymetrix. The test design, experimentation, data collection, analysis, and interpretation were done independently by the researchers. ![]()

Credit: NIGMS
NASHVILLE—New research suggests we may need to use more sensitive methods to analyze patients with cytogenetically normal acute myeloid leukemia or myelodysplastic syndrome (CN-AML/MDS).
Using “highly sensitive” microarray technology, researchers found a distinct pattern of genetic abnormalities in 22 patients diagnosed with CN-AML/MDS.
The team identified 3 overlapping regions of homozygosity in 3 genes, 2 of which are known to be involved in carcinogenesis.
This suggests that using karyotyping or FISH, or simply looking for known mutations, is not sufficient for evaluating patients with CN-AML/MDS, according to Ravindra Kolhe, MD, PhD, of the Medical College of Georgia at Georgia Regents University.
“The technology we currently use can’t identify specifically what’s wrong,” Dr Kolhe said. “We have to use more sensitive tests to give patients the proper answer.”
Dr Kolhe presented this finding, and the research to support it, at the American College of Medical Genetics and Genomics Annual Clinical Genetics Meeting.
He and his colleagues analyzed 22 patients. Seventeen had AML, and 5 had MDS, including 1 with refractory anemia with excess blasts-2. All patients had normal karyotype and FISH and had greater than 20% blasts in the bone marrow.
The researchers analyzed samples from these patients using a high-resolution, single-nucleotide polymorphism (SNP) microarray called CytoScanHD.
According to the company that markets this technology (Affymetrix, Inc.), the assay includes 750,000 SNPs with over 99% accuracy to detect accurate breakpoint estimation, loss of heterozygosity determination, regions identical-by-descent, maternal contamination, and low-level mosaicism.
For Dr Kolhe and his colleagues, the assay revealed small, previously undetectable changes in patients thought to be cytogenetically normal.
Specifically, the researchers identified 3 overlapping regions of homozygosity in all 22 cases—chromosome 1p34.3, chromosome 1p32.3, and chromosome 16q22.1 in the SFPQ, EPS15, and CTCF genes, respectively.
SFPQ and CTCF are already known to be involved in carcinogenesis, and Dr Kolhe and his colleagues are now investigating the role of EPS15 in leukemogenesis.
The researchers also identified additional abnormalities and are investigating these as well. They are sequencing the genes to identify homozygous or compound heterozygous mutations, performing expression studies to confirm that these mutations are leukemic, and conducting experiments in knockout mice to demonstrate that these genes produce the same leukemia phenotype.
The materials and reagents for this study were provided by Affymetrix. The test design, experimentation, data collection, analysis, and interpretation were done independently by the researchers. ![]()

Credit: NIGMS
NASHVILLE—New research suggests we may need to use more sensitive methods to analyze patients with cytogenetically normal acute myeloid leukemia or myelodysplastic syndrome (CN-AML/MDS).
Using “highly sensitive” microarray technology, researchers found a distinct pattern of genetic abnormalities in 22 patients diagnosed with CN-AML/MDS.
The team identified 3 overlapping regions of homozygosity in 3 genes, 2 of which are known to be involved in carcinogenesis.
This suggests that using karyotyping or FISH, or simply looking for known mutations, is not sufficient for evaluating patients with CN-AML/MDS, according to Ravindra Kolhe, MD, PhD, of the Medical College of Georgia at Georgia Regents University.
“The technology we currently use can’t identify specifically what’s wrong,” Dr Kolhe said. “We have to use more sensitive tests to give patients the proper answer.”
Dr Kolhe presented this finding, and the research to support it, at the American College of Medical Genetics and Genomics Annual Clinical Genetics Meeting.
He and his colleagues analyzed 22 patients. Seventeen had AML, and 5 had MDS, including 1 with refractory anemia with excess blasts-2. All patients had normal karyotype and FISH and had greater than 20% blasts in the bone marrow.
The researchers analyzed samples from these patients using a high-resolution, single-nucleotide polymorphism (SNP) microarray called CytoScanHD.
According to the company that markets this technology (Affymetrix, Inc.), the assay includes 750,000 SNPs with over 99% accuracy to detect accurate breakpoint estimation, loss of heterozygosity determination, regions identical-by-descent, maternal contamination, and low-level mosaicism.
For Dr Kolhe and his colleagues, the assay revealed small, previously undetectable changes in patients thought to be cytogenetically normal.
Specifically, the researchers identified 3 overlapping regions of homozygosity in all 22 cases—chromosome 1p34.3, chromosome 1p32.3, and chromosome 16q22.1 in the SFPQ, EPS15, and CTCF genes, respectively.
SFPQ and CTCF are already known to be involved in carcinogenesis, and Dr Kolhe and his colleagues are now investigating the role of EPS15 in leukemogenesis.
The researchers also identified additional abnormalities and are investigating these as well. They are sequencing the genes to identify homozygous or compound heterozygous mutations, performing expression studies to confirm that these mutations are leukemic, and conducting experiments in knockout mice to demonstrate that these genes produce the same leukemia phenotype.
The materials and reagents for this study were provided by Affymetrix. The test design, experimentation, data collection, analysis, and interpretation were done independently by the researchers. ![]()
Monosomal karyotype, high prognostic risk score predicted transplantation failure
Monosomal karyotype and high prognostic risk according to the revised International Prognostic Scoring System are independent predictors of relapse and mortality in patients with myelodysplastic syndrome or oligoblastic acute myeloid leukemia who undergo allogeneic hematopoietic stem cell transplantation, according to findings from the GITMO (Gruppo Italiano Trapianto di Midollo Osseo) registry.
Treatment failure after allogeneic hematopoietic stem cell transplantation may be from transplant complications or relapse. To understand the predictors of failure, investigators studied outcomes in 519 patients with myelodysplastic syndrome or oligoblastic acute myeloid leukemia who underwent hematopoietic stem cell transplantation between 2000 and 2011.
Those with monosomal karyotype had a 49% relapse rate and a 10% 5-year overall survival rate; both rates were significantly worse, compared with patients without monosomal karyotype (P less than .001 for each). Those considered high or very-high risk based on the International Prognostic Scoring System (IPSS-R), had 39% and 23% 5-year overall survival, respectively, and 23% and 39% relapse rates, respectively (P less than .001 in all cases vs. patients not at high or very-high risk), Dr. Matteo G. Della Porta of Fondazione IRCCS Policlinico San Matteo, Pavia, Italy and colleagues reported on behalf of the GITMO.
Age of 50 years or older and high hematopoietic cell transplantation-comorbidity index scores were independent predictors of nonrelapse mortality (P = .02; P = .017, respectively), they found (Blood 2014 [doi:10.1182/blood-2013-12-542720]).
Accounting for various combinations of patients’ ages, IPSS-R category, monosomal karyotype, and high hematopoietic cell transplantation–comorbidity index, the 5-year probability of survival after allogeneic hematopoietic stem cell transplantation ranged from 0 to 94%. The analyses performed reinforce the concept that allogenic hematopoietic stem cell transplantation – the only potentially curative treatment for MDS – "offers optimal eradication of myelodysplastic hematopoiesis when the procedure is performed before MDS patients progress to advanced disease stages," the investigators concluded.
The investigators reported having no disclosures.
Monosomal karyotype and high prognostic risk according to the revised International Prognostic Scoring System are independent predictors of relapse and mortality in patients with myelodysplastic syndrome or oligoblastic acute myeloid leukemia who undergo allogeneic hematopoietic stem cell transplantation, according to findings from the GITMO (Gruppo Italiano Trapianto di Midollo Osseo) registry.
Treatment failure after allogeneic hematopoietic stem cell transplantation may be from transplant complications or relapse. To understand the predictors of failure, investigators studied outcomes in 519 patients with myelodysplastic syndrome or oligoblastic acute myeloid leukemia who underwent hematopoietic stem cell transplantation between 2000 and 2011.
Those with monosomal karyotype had a 49% relapse rate and a 10% 5-year overall survival rate; both rates were significantly worse, compared with patients without monosomal karyotype (P less than .001 for each). Those considered high or very-high risk based on the International Prognostic Scoring System (IPSS-R), had 39% and 23% 5-year overall survival, respectively, and 23% and 39% relapse rates, respectively (P less than .001 in all cases vs. patients not at high or very-high risk), Dr. Matteo G. Della Porta of Fondazione IRCCS Policlinico San Matteo, Pavia, Italy and colleagues reported on behalf of the GITMO.
Age of 50 years or older and high hematopoietic cell transplantation-comorbidity index scores were independent predictors of nonrelapse mortality (P = .02; P = .017, respectively), they found (Blood 2014 [doi:10.1182/blood-2013-12-542720]).
Accounting for various combinations of patients’ ages, IPSS-R category, monosomal karyotype, and high hematopoietic cell transplantation–comorbidity index, the 5-year probability of survival after allogeneic hematopoietic stem cell transplantation ranged from 0 to 94%. The analyses performed reinforce the concept that allogenic hematopoietic stem cell transplantation – the only potentially curative treatment for MDS – "offers optimal eradication of myelodysplastic hematopoiesis when the procedure is performed before MDS patients progress to advanced disease stages," the investigators concluded.
The investigators reported having no disclosures.
Monosomal karyotype and high prognostic risk according to the revised International Prognostic Scoring System are independent predictors of relapse and mortality in patients with myelodysplastic syndrome or oligoblastic acute myeloid leukemia who undergo allogeneic hematopoietic stem cell transplantation, according to findings from the GITMO (Gruppo Italiano Trapianto di Midollo Osseo) registry.
Treatment failure after allogeneic hematopoietic stem cell transplantation may be from transplant complications or relapse. To understand the predictors of failure, investigators studied outcomes in 519 patients with myelodysplastic syndrome or oligoblastic acute myeloid leukemia who underwent hematopoietic stem cell transplantation between 2000 and 2011.
Those with monosomal karyotype had a 49% relapse rate and a 10% 5-year overall survival rate; both rates were significantly worse, compared with patients without monosomal karyotype (P less than .001 for each). Those considered high or very-high risk based on the International Prognostic Scoring System (IPSS-R), had 39% and 23% 5-year overall survival, respectively, and 23% and 39% relapse rates, respectively (P less than .001 in all cases vs. patients not at high or very-high risk), Dr. Matteo G. Della Porta of Fondazione IRCCS Policlinico San Matteo, Pavia, Italy and colleagues reported on behalf of the GITMO.
Age of 50 years or older and high hematopoietic cell transplantation-comorbidity index scores were independent predictors of nonrelapse mortality (P = .02; P = .017, respectively), they found (Blood 2014 [doi:10.1182/blood-2013-12-542720]).
Accounting for various combinations of patients’ ages, IPSS-R category, monosomal karyotype, and high hematopoietic cell transplantation–comorbidity index, the 5-year probability of survival after allogeneic hematopoietic stem cell transplantation ranged from 0 to 94%. The analyses performed reinforce the concept that allogenic hematopoietic stem cell transplantation – the only potentially curative treatment for MDS – "offers optimal eradication of myelodysplastic hematopoiesis when the procedure is performed before MDS patients progress to advanced disease stages," the investigators concluded.
The investigators reported having no disclosures.
FROM BLOOD
Major finding: Patients with a monosomal karyotype had a 49% relapse rate and a 10% 5-year overall survival rate, and those considered high or very-high risk based on the IPSS-R, had 39% and 23% 5-year overall survival, respectively, and 23% and 39% relapse rates, respectively.
Data source: An analysis of GITMO registry data.
Disclosures: The investigators reported having no disclosures.
Group grows functional LSCs in culture
Two small-molecule compounds can help researchers maintain leukemic stem cells (LSCs) in culture, according to a paper published in Nature Methods.
Investigators said they created improved culture conditions for primary human acute myeloid leukemia (AML) cells, based on serum-free medium supplemented with the small molecules SR1 and UM729.
These conditions increased the yield of phenotypically undifferentiated CD34+ AML cells and supported the ex vivo maintenance of LSCs that are typically lost in culture.
Caroline Pabst, MD, of the Institute for Research in Immunology and Cancer at the University of Montreal in Quebec, Canada, and her colleagues conducted this research using AML patient samples.
The team screened about 6000 compounds in an attempt to identify small molecules that promote the ex vivo expansion of undifferentiated AML cells.
And they found that suppressors of the aryl-hydrocarbon receptor (AhR) pathway were enriched among the hit compounds.
So the researchers decided to study 2 chemically distinct AhR suppressors: N-methyl-β-carboline-3-carboxamide (C05), which yielded the highest CD34+CD15- cell counts in secondary screens, and the known AhR antagonist SR1. They also studied the pyrimidoindole UM729, which had shown no effects on AhR target genes.
Experiments showed that the AhR pathway was “rapidly and robustly” activated in the AML samples upon culture. However, suppressing the pathway with SR1 and C05 enabled the expansion of CD34+ AML cells and supported the maintenance of LSCs.
In addition, UM729 had an additive effect with SR1 on the maintenance of AML stem and progenitor cells in vitro.
The investigators said these results should help establish defined conditions to overcome spontaneous differentiation and cell death in ex vivo cultures of primary human AML specimens.
The team believes at least 3 molecular targets could be involved in this process, and 2 of them are targeted by SR1 and UM729.
So these compounds could serve as a standardized supplement to culture media. They might aid studies of self-renewal mechanisms and help researchers identify new antileukemic drugs. ![]()

Two small-molecule compounds can help researchers maintain leukemic stem cells (LSCs) in culture, according to a paper published in Nature Methods.
Investigators said they created improved culture conditions for primary human acute myeloid leukemia (AML) cells, based on serum-free medium supplemented with the small molecules SR1 and UM729.
These conditions increased the yield of phenotypically undifferentiated CD34+ AML cells and supported the ex vivo maintenance of LSCs that are typically lost in culture.
Caroline Pabst, MD, of the Institute for Research in Immunology and Cancer at the University of Montreal in Quebec, Canada, and her colleagues conducted this research using AML patient samples.
The team screened about 6000 compounds in an attempt to identify small molecules that promote the ex vivo expansion of undifferentiated AML cells.
And they found that suppressors of the aryl-hydrocarbon receptor (AhR) pathway were enriched among the hit compounds.
So the researchers decided to study 2 chemically distinct AhR suppressors: N-methyl-β-carboline-3-carboxamide (C05), which yielded the highest CD34+CD15- cell counts in secondary screens, and the known AhR antagonist SR1. They also studied the pyrimidoindole UM729, which had shown no effects on AhR target genes.
Experiments showed that the AhR pathway was “rapidly and robustly” activated in the AML samples upon culture. However, suppressing the pathway with SR1 and C05 enabled the expansion of CD34+ AML cells and supported the maintenance of LSCs.
In addition, UM729 had an additive effect with SR1 on the maintenance of AML stem and progenitor cells in vitro.
The investigators said these results should help establish defined conditions to overcome spontaneous differentiation and cell death in ex vivo cultures of primary human AML specimens.
The team believes at least 3 molecular targets could be involved in this process, and 2 of them are targeted by SR1 and UM729.
So these compounds could serve as a standardized supplement to culture media. They might aid studies of self-renewal mechanisms and help researchers identify new antileukemic drugs. ![]()

Two small-molecule compounds can help researchers maintain leukemic stem cells (LSCs) in culture, according to a paper published in Nature Methods.
Investigators said they created improved culture conditions for primary human acute myeloid leukemia (AML) cells, based on serum-free medium supplemented with the small molecules SR1 and UM729.
These conditions increased the yield of phenotypically undifferentiated CD34+ AML cells and supported the ex vivo maintenance of LSCs that are typically lost in culture.
Caroline Pabst, MD, of the Institute for Research in Immunology and Cancer at the University of Montreal in Quebec, Canada, and her colleagues conducted this research using AML patient samples.
The team screened about 6000 compounds in an attempt to identify small molecules that promote the ex vivo expansion of undifferentiated AML cells.
And they found that suppressors of the aryl-hydrocarbon receptor (AhR) pathway were enriched among the hit compounds.
So the researchers decided to study 2 chemically distinct AhR suppressors: N-methyl-β-carboline-3-carboxamide (C05), which yielded the highest CD34+CD15- cell counts in secondary screens, and the known AhR antagonist SR1. They also studied the pyrimidoindole UM729, which had shown no effects on AhR target genes.
Experiments showed that the AhR pathway was “rapidly and robustly” activated in the AML samples upon culture. However, suppressing the pathway with SR1 and C05 enabled the expansion of CD34+ AML cells and supported the maintenance of LSCs.
In addition, UM729 had an additive effect with SR1 on the maintenance of AML stem and progenitor cells in vitro.
The investigators said these results should help establish defined conditions to overcome spontaneous differentiation and cell death in ex vivo cultures of primary human AML specimens.
The team believes at least 3 molecular targets could be involved in this process, and 2 of them are targeted by SR1 and UM729.
So these compounds could serve as a standardized supplement to culture media. They might aid studies of self-renewal mechanisms and help researchers identify new antileukemic drugs. ![]()
CNS involvement doesn’t affect survival after allo-SCT
GRAPEVINE, TEXAS—Results of a large, retrospective study suggest that allogeneic stem cell transplant (allo-SCT) can overcome the poor prognosis associated with central nervous system (CNS) involvement in acute myeloid leukemia (AML).
By analyzing transplant outcomes in more than 5000 patients, researchers found that subjects with CNS AML had rates of relapse and survival that were similar to those of patients without CNS involvement.
The team also identified factors that can predict for survival in CNS AML, including cytogenetic risk group, the presence of chronic GVHD, and whether a patient was in complete response at transplant.
Jun Aoki, MD, of Tokyo Metropolitan Komagome Hospital in Japan, presented these findings at the 2014 BMT Tandem Meetings as abstract 68.
Dr Aoki pointed out that CNS involvement is rare in adult AML, occurring in about 5% of patients. However, these patients generally have poor prognosis. And although allo-SCT is one of the options used to treat CNS AML, exactly how CNS involvement impacts transplant outcomes remains unclear.
So Dr Aoki and his colleagues conducted a nationwide, retrospective study to gain some insight. They collected data from the registry database of the Japan Society for Hematopoietic Cell Transplantation.
Patients had to be older than 15 years of age, have their first allo-SCT between 2006 and 2011, and not have acute promyelocytic leukemia.
The researchers identified 5068 patients who met these criteria, and 157 of them had CNS AML. CNS involvement was defined as infiltration of leukemia cells into CNS or myeloid sarcoma in CNS that were identified at any time from diagnosis to transplant.
No difference in relapse, survival
There were no significant differences between CNS patients and controls with regard to the estimated overall survival (OS), leukemia-free survival, cumulative incidence of relapse, or non-relapse mortality at 5 years.
OS was 39.9% among controls and 38.5% among CNS patients (P=0.847). Leukemia-free survival was 41.2% and 41.5%, respectively (P=0.82).
The cumulative incidence of relapse was 29.8% among controls and 31.8% among CNS patients (P=0.418). And non-relapse mortality was 22.5% and 26.5%, respectively (P=0.142).
Factors predicting OS
To determine the impact of patient and treatment characteristics on OS, the researchers conducted a multivariate analysis. This confirmed that CNS involvement was not a risk factor for OS.
But it revealed a number of other factors that adversely affect OS, including age of 50 or older (P<0.001), lack of a complete response at allo-SCT (P<0.001), a donor source of unrelated cord blood (P=0.005), having a prognostic score of 2-4 (P<0.001), unfavorable cytogenetics (P<0.001), and the absence of acute or chronic GVHD (P<0.001 for both).
When the researchers analyzed only CNS patients, they discovered that not all of these factors retained significance. Only the absence of chronic GVHD (P=0.002), lack of complete response at transplant (P<0.001), and having either intermediate (P=0.003) or unfavorable cytogenetics (P=0.011) were adversely associated with OS in these patients. ![]()
GRAPEVINE, TEXAS—Results of a large, retrospective study suggest that allogeneic stem cell transplant (allo-SCT) can overcome the poor prognosis associated with central nervous system (CNS) involvement in acute myeloid leukemia (AML).
By analyzing transplant outcomes in more than 5000 patients, researchers found that subjects with CNS AML had rates of relapse and survival that were similar to those of patients without CNS involvement.
The team also identified factors that can predict for survival in CNS AML, including cytogenetic risk group, the presence of chronic GVHD, and whether a patient was in complete response at transplant.
Jun Aoki, MD, of Tokyo Metropolitan Komagome Hospital in Japan, presented these findings at the 2014 BMT Tandem Meetings as abstract 68.
Dr Aoki pointed out that CNS involvement is rare in adult AML, occurring in about 5% of patients. However, these patients generally have poor prognosis. And although allo-SCT is one of the options used to treat CNS AML, exactly how CNS involvement impacts transplant outcomes remains unclear.
So Dr Aoki and his colleagues conducted a nationwide, retrospective study to gain some insight. They collected data from the registry database of the Japan Society for Hematopoietic Cell Transplantation.
Patients had to be older than 15 years of age, have their first allo-SCT between 2006 and 2011, and not have acute promyelocytic leukemia.
The researchers identified 5068 patients who met these criteria, and 157 of them had CNS AML. CNS involvement was defined as infiltration of leukemia cells into CNS or myeloid sarcoma in CNS that were identified at any time from diagnosis to transplant.
No difference in relapse, survival
There were no significant differences between CNS patients and controls with regard to the estimated overall survival (OS), leukemia-free survival, cumulative incidence of relapse, or non-relapse mortality at 5 years.
OS was 39.9% among controls and 38.5% among CNS patients (P=0.847). Leukemia-free survival was 41.2% and 41.5%, respectively (P=0.82).
The cumulative incidence of relapse was 29.8% among controls and 31.8% among CNS patients (P=0.418). And non-relapse mortality was 22.5% and 26.5%, respectively (P=0.142).
Factors predicting OS
To determine the impact of patient and treatment characteristics on OS, the researchers conducted a multivariate analysis. This confirmed that CNS involvement was not a risk factor for OS.
But it revealed a number of other factors that adversely affect OS, including age of 50 or older (P<0.001), lack of a complete response at allo-SCT (P<0.001), a donor source of unrelated cord blood (P=0.005), having a prognostic score of 2-4 (P<0.001), unfavorable cytogenetics (P<0.001), and the absence of acute or chronic GVHD (P<0.001 for both).
When the researchers analyzed only CNS patients, they discovered that not all of these factors retained significance. Only the absence of chronic GVHD (P=0.002), lack of complete response at transplant (P<0.001), and having either intermediate (P=0.003) or unfavorable cytogenetics (P=0.011) were adversely associated with OS in these patients. ![]()
GRAPEVINE, TEXAS—Results of a large, retrospective study suggest that allogeneic stem cell transplant (allo-SCT) can overcome the poor prognosis associated with central nervous system (CNS) involvement in acute myeloid leukemia (AML).
By analyzing transplant outcomes in more than 5000 patients, researchers found that subjects with CNS AML had rates of relapse and survival that were similar to those of patients without CNS involvement.
The team also identified factors that can predict for survival in CNS AML, including cytogenetic risk group, the presence of chronic GVHD, and whether a patient was in complete response at transplant.
Jun Aoki, MD, of Tokyo Metropolitan Komagome Hospital in Japan, presented these findings at the 2014 BMT Tandem Meetings as abstract 68.
Dr Aoki pointed out that CNS involvement is rare in adult AML, occurring in about 5% of patients. However, these patients generally have poor prognosis. And although allo-SCT is one of the options used to treat CNS AML, exactly how CNS involvement impacts transplant outcomes remains unclear.
So Dr Aoki and his colleagues conducted a nationwide, retrospective study to gain some insight. They collected data from the registry database of the Japan Society for Hematopoietic Cell Transplantation.
Patients had to be older than 15 years of age, have their first allo-SCT between 2006 and 2011, and not have acute promyelocytic leukemia.
The researchers identified 5068 patients who met these criteria, and 157 of them had CNS AML. CNS involvement was defined as infiltration of leukemia cells into CNS or myeloid sarcoma in CNS that were identified at any time from diagnosis to transplant.
No difference in relapse, survival
There were no significant differences between CNS patients and controls with regard to the estimated overall survival (OS), leukemia-free survival, cumulative incidence of relapse, or non-relapse mortality at 5 years.
OS was 39.9% among controls and 38.5% among CNS patients (P=0.847). Leukemia-free survival was 41.2% and 41.5%, respectively (P=0.82).
The cumulative incidence of relapse was 29.8% among controls and 31.8% among CNS patients (P=0.418). And non-relapse mortality was 22.5% and 26.5%, respectively (P=0.142).
Factors predicting OS
To determine the impact of patient and treatment characteristics on OS, the researchers conducted a multivariate analysis. This confirmed that CNS involvement was not a risk factor for OS.
But it revealed a number of other factors that adversely affect OS, including age of 50 or older (P<0.001), lack of a complete response at allo-SCT (P<0.001), a donor source of unrelated cord blood (P=0.005), having a prognostic score of 2-4 (P<0.001), unfavorable cytogenetics (P<0.001), and the absence of acute or chronic GVHD (P<0.001 for both).
When the researchers analyzed only CNS patients, they discovered that not all of these factors retained significance. Only the absence of chronic GVHD (P=0.002), lack of complete response at transplant (P<0.001), and having either intermediate (P=0.003) or unfavorable cytogenetics (P=0.011) were adversely associated with OS in these patients. ![]()
Drugs get orphan designation for AML, MM

Credit: FDA
The US Food and Drug Administration (FDA) has granted orphan designation for the histone deacetylase (HDAC) inhibitor pracinostat to treat acute myeloid leukemia (AML) and the proteasome inhibitor marizomib to treat multiple myeloma (MM).
Orphan designation is available for drugs that treat or prevent rare diseases affecting fewer than 200,000 people in the US.
The designation qualifies the sponsor of a drug for development incentives, including tax credits for qualified clinical testing, prescription drug user fee exemptions, and 7-year marketing exclusivity upon FDA approval.
About pracinostat
The oral HDAC inhibitor pracinostat has been tested in phase 1 and 2 trials of adult and pediatric patients with advanced hematologic disorders and solid tumors.
The drug has been generally well tolerated in more than 200 patients to date, according to the drug’s maker, MEI Pharma.
In a dose-escalation phase 1 trial, pracinostat demonstrated single-agent activity in elderly AML patients. Two of 14 patients (14%) achieved a complete remission, with responses persisting more than 206 days and 362 days.
Researchers are currently conducting a phase 2 trial of pracinostat in combination with azacitidine in elderly patients with newly diagnosed AML. Preliminary data from this trial are expected to be available by December 2014.
About marizomib
The proteasome inhibitor marizomib is under development for the treatment of MM and other malignancies.
Intravenous marizomib has been evaluated in more than 230 patients across 4 phase 1/2 studies, as a single agent or in combination with dexamethasone or an HDAC inhibitor.
Researchers are currently evaluating marizomib in combination with dexamethasone in an ongoing phase 2 trial of highly refractory MM patients, including those who are refractory to carfilzomib.
Marizomib is also being tested in combination with pomalidomide and dexamethasone in a phase 1/2 study of patients with relapsed and refractory MM.
The drug’s maker, Triphase Accelerator Corporation, is currently developing an oral formulation of marizomib. ![]()

Credit: FDA
The US Food and Drug Administration (FDA) has granted orphan designation for the histone deacetylase (HDAC) inhibitor pracinostat to treat acute myeloid leukemia (AML) and the proteasome inhibitor marizomib to treat multiple myeloma (MM).
Orphan designation is available for drugs that treat or prevent rare diseases affecting fewer than 200,000 people in the US.
The designation qualifies the sponsor of a drug for development incentives, including tax credits for qualified clinical testing, prescription drug user fee exemptions, and 7-year marketing exclusivity upon FDA approval.
About pracinostat
The oral HDAC inhibitor pracinostat has been tested in phase 1 and 2 trials of adult and pediatric patients with advanced hematologic disorders and solid tumors.
The drug has been generally well tolerated in more than 200 patients to date, according to the drug’s maker, MEI Pharma.
In a dose-escalation phase 1 trial, pracinostat demonstrated single-agent activity in elderly AML patients. Two of 14 patients (14%) achieved a complete remission, with responses persisting more than 206 days and 362 days.
Researchers are currently conducting a phase 2 trial of pracinostat in combination with azacitidine in elderly patients with newly diagnosed AML. Preliminary data from this trial are expected to be available by December 2014.
About marizomib
The proteasome inhibitor marizomib is under development for the treatment of MM and other malignancies.
Intravenous marizomib has been evaluated in more than 230 patients across 4 phase 1/2 studies, as a single agent or in combination with dexamethasone or an HDAC inhibitor.
Researchers are currently evaluating marizomib in combination with dexamethasone in an ongoing phase 2 trial of highly refractory MM patients, including those who are refractory to carfilzomib.
Marizomib is also being tested in combination with pomalidomide and dexamethasone in a phase 1/2 study of patients with relapsed and refractory MM.
The drug’s maker, Triphase Accelerator Corporation, is currently developing an oral formulation of marizomib. ![]()

Credit: FDA
The US Food and Drug Administration (FDA) has granted orphan designation for the histone deacetylase (HDAC) inhibitor pracinostat to treat acute myeloid leukemia (AML) and the proteasome inhibitor marizomib to treat multiple myeloma (MM).
Orphan designation is available for drugs that treat or prevent rare diseases affecting fewer than 200,000 people in the US.
The designation qualifies the sponsor of a drug for development incentives, including tax credits for qualified clinical testing, prescription drug user fee exemptions, and 7-year marketing exclusivity upon FDA approval.
About pracinostat
The oral HDAC inhibitor pracinostat has been tested in phase 1 and 2 trials of adult and pediatric patients with advanced hematologic disorders and solid tumors.
The drug has been generally well tolerated in more than 200 patients to date, according to the drug’s maker, MEI Pharma.
In a dose-escalation phase 1 trial, pracinostat demonstrated single-agent activity in elderly AML patients. Two of 14 patients (14%) achieved a complete remission, with responses persisting more than 206 days and 362 days.
Researchers are currently conducting a phase 2 trial of pracinostat in combination with azacitidine in elderly patients with newly diagnosed AML. Preliminary data from this trial are expected to be available by December 2014.
About marizomib
The proteasome inhibitor marizomib is under development for the treatment of MM and other malignancies.
Intravenous marizomib has been evaluated in more than 230 patients across 4 phase 1/2 studies, as a single agent or in combination with dexamethasone or an HDAC inhibitor.
Researchers are currently evaluating marizomib in combination with dexamethasone in an ongoing phase 2 trial of highly refractory MM patients, including those who are refractory to carfilzomib.
Marizomib is also being tested in combination with pomalidomide and dexamethasone in a phase 1/2 study of patients with relapsed and refractory MM.
The drug’s maker, Triphase Accelerator Corporation, is currently developing an oral formulation of marizomib. ![]()
Investigating the cause of infant leukemias
Infants who develop leukemia during the first year of life inherit a combination of genetic variations that can make them highly susceptible to the disease, according to a study published in Leukemia.
Results of whole-exome sequencing suggested that infants with leukemia inherited genetic variants from both parents that, by themselves, would not cause leukemia but, in combination, put the infants at high risk of developing the disease.
“We sequenced every single gene and found that infants with leukemia were born with an excess of damaging changes in genes known to be linked to leukemia,” said study author Todd Druley, MD, PhD, of Washington University School of Medicine in St Louis, Missouri.
“For each child, both parents carried a few harmful genetic variations in their DNA, and, just by chance, their child inherited all of these changes.”
However, it’s unlikely that the inherited variations alone cause leukemia, Dr Druley said. The infants likely needed to accumulate a few additional variations.
To uncover these findings, Dr Druley and his colleagues performed whole-exome sequencing in infants with acute myeloid leukemia (AML), infants with acute lymphoblastic leukemia (ALL), and the mothers of these children. The researchers used the process of elimination to determine a father’s contribution to a child’s DNA.
Among the 23 families studied, there was no history of pediatric cancers. As a comparison, the researchers also sequenced the DNA of 25 healthy children.
The team found the average amount of congenital coding variations was higher in infants with leukemia than in their mothers or the control subjects. The average total variants per exome was 1264.4 in infants with ALL, 1112.6 in their mothers, 2549.9 in infants with AML, 1225.0 in their mothers, and 582.8 in healthy controls.
The researchers then decided to home in on variants that were likely to impart a functional effect associated with leukemia. Using the COSMIC database, the team identified 126 ALL-associated genes and 655 AML-associated genes.
They found an average of 12.1 variants per ALL patient in the ALL-associated genes and 163.4 variants per AML patient in the AML-associated genes. There were 6.4 ALL-associated variants in the ALL patients’ mothers, 132.5 AML-associated variants in the AML patients’ mothers, 1.9 ALL variants in controls, and 27.5 AML variants in controls.
To prioritize genes that might be most relevant to infant leukemia, the researchers looked for compound heterozygous genes and the genes that were most commonly variant in all patients.
All of the infants with AML and 50% of the infants with ALL were compound heterozygotes for MLL3. Sixty-seven percent of AML patients were compound heterozygotes for RYR1 and FLG, and 50% of ALL patients were compound heterozygotes for RBMX.
The most variant (but not necessarily compound heterozygous) AML-associated genes in infants with AML were TTN, MLL3, and FLG. But the ALL-associated genes MDN1, SYNE1, and MLL2 were frequently variable in AML patients as well.
For infants with ALL, MDN1 was the most variable ALL-associated gene. But these infants also had frequent variations in the AML-associated genes TTN, RBMX, and MLL3.
Dr Druley and his colleagues plan to study these variations in more detail to understand how they contribute to infant leukemia development. ![]()
Infants who develop leukemia during the first year of life inherit a combination of genetic variations that can make them highly susceptible to the disease, according to a study published in Leukemia.
Results of whole-exome sequencing suggested that infants with leukemia inherited genetic variants from both parents that, by themselves, would not cause leukemia but, in combination, put the infants at high risk of developing the disease.
“We sequenced every single gene and found that infants with leukemia were born with an excess of damaging changes in genes known to be linked to leukemia,” said study author Todd Druley, MD, PhD, of Washington University School of Medicine in St Louis, Missouri.
“For each child, both parents carried a few harmful genetic variations in their DNA, and, just by chance, their child inherited all of these changes.”
However, it’s unlikely that the inherited variations alone cause leukemia, Dr Druley said. The infants likely needed to accumulate a few additional variations.
To uncover these findings, Dr Druley and his colleagues performed whole-exome sequencing in infants with acute myeloid leukemia (AML), infants with acute lymphoblastic leukemia (ALL), and the mothers of these children. The researchers used the process of elimination to determine a father’s contribution to a child’s DNA.
Among the 23 families studied, there was no history of pediatric cancers. As a comparison, the researchers also sequenced the DNA of 25 healthy children.
The team found the average amount of congenital coding variations was higher in infants with leukemia than in their mothers or the control subjects. The average total variants per exome was 1264.4 in infants with ALL, 1112.6 in their mothers, 2549.9 in infants with AML, 1225.0 in their mothers, and 582.8 in healthy controls.
The researchers then decided to home in on variants that were likely to impart a functional effect associated with leukemia. Using the COSMIC database, the team identified 126 ALL-associated genes and 655 AML-associated genes.
They found an average of 12.1 variants per ALL patient in the ALL-associated genes and 163.4 variants per AML patient in the AML-associated genes. There were 6.4 ALL-associated variants in the ALL patients’ mothers, 132.5 AML-associated variants in the AML patients’ mothers, 1.9 ALL variants in controls, and 27.5 AML variants in controls.
To prioritize genes that might be most relevant to infant leukemia, the researchers looked for compound heterozygous genes and the genes that were most commonly variant in all patients.
All of the infants with AML and 50% of the infants with ALL were compound heterozygotes for MLL3. Sixty-seven percent of AML patients were compound heterozygotes for RYR1 and FLG, and 50% of ALL patients were compound heterozygotes for RBMX.
The most variant (but not necessarily compound heterozygous) AML-associated genes in infants with AML were TTN, MLL3, and FLG. But the ALL-associated genes MDN1, SYNE1, and MLL2 were frequently variable in AML patients as well.
For infants with ALL, MDN1 was the most variable ALL-associated gene. But these infants also had frequent variations in the AML-associated genes TTN, RBMX, and MLL3.
Dr Druley and his colleagues plan to study these variations in more detail to understand how they contribute to infant leukemia development. ![]()
Infants who develop leukemia during the first year of life inherit a combination of genetic variations that can make them highly susceptible to the disease, according to a study published in Leukemia.
Results of whole-exome sequencing suggested that infants with leukemia inherited genetic variants from both parents that, by themselves, would not cause leukemia but, in combination, put the infants at high risk of developing the disease.
“We sequenced every single gene and found that infants with leukemia were born with an excess of damaging changes in genes known to be linked to leukemia,” said study author Todd Druley, MD, PhD, of Washington University School of Medicine in St Louis, Missouri.
“For each child, both parents carried a few harmful genetic variations in their DNA, and, just by chance, their child inherited all of these changes.”
However, it’s unlikely that the inherited variations alone cause leukemia, Dr Druley said. The infants likely needed to accumulate a few additional variations.
To uncover these findings, Dr Druley and his colleagues performed whole-exome sequencing in infants with acute myeloid leukemia (AML), infants with acute lymphoblastic leukemia (ALL), and the mothers of these children. The researchers used the process of elimination to determine a father’s contribution to a child’s DNA.
Among the 23 families studied, there was no history of pediatric cancers. As a comparison, the researchers also sequenced the DNA of 25 healthy children.
The team found the average amount of congenital coding variations was higher in infants with leukemia than in their mothers or the control subjects. The average total variants per exome was 1264.4 in infants with ALL, 1112.6 in their mothers, 2549.9 in infants with AML, 1225.0 in their mothers, and 582.8 in healthy controls.
The researchers then decided to home in on variants that were likely to impart a functional effect associated with leukemia. Using the COSMIC database, the team identified 126 ALL-associated genes and 655 AML-associated genes.
They found an average of 12.1 variants per ALL patient in the ALL-associated genes and 163.4 variants per AML patient in the AML-associated genes. There were 6.4 ALL-associated variants in the ALL patients’ mothers, 132.5 AML-associated variants in the AML patients’ mothers, 1.9 ALL variants in controls, and 27.5 AML variants in controls.
To prioritize genes that might be most relevant to infant leukemia, the researchers looked for compound heterozygous genes and the genes that were most commonly variant in all patients.
All of the infants with AML and 50% of the infants with ALL were compound heterozygotes for MLL3. Sixty-seven percent of AML patients were compound heterozygotes for RYR1 and FLG, and 50% of ALL patients were compound heterozygotes for RBMX.
The most variant (but not necessarily compound heterozygous) AML-associated genes in infants with AML were TTN, MLL3, and FLG. But the ALL-associated genes MDN1, SYNE1, and MLL2 were frequently variable in AML patients as well.
For infants with ALL, MDN1 was the most variable ALL-associated gene. But these infants also had frequent variations in the AML-associated genes TTN, RBMX, and MLL3.
Dr Druley and his colleagues plan to study these variations in more detail to understand how they contribute to infant leukemia development. ![]()
Protein may be target for AML treatment

Credit: Rhoda Baer
The protein WTAP could play an important role in the development of acute myeloid leukemia (AML), according to new research.
Investigators discovered that AML cells have higher-than-normal levels of WTAP.
But silencing WTAP expression in leukemic cells can suppress proliferation and induce differentiation.
And, in mouse models of AML, knocking down WTAP can reduce tumor growth.
The researchers recounted these findings in a letter to Leukemia.
The team first uncovered high levels of WTAP in AML cells compared to normal peripheral blood mononuclear cells. And they found evidence to suggest that this contributes to abnormal cell behavior.
WTAP levels were not associated with individual cytogenetic abnormalities, but FLT3-ITD and NPM1 mutations were significantly correlated with WTAP expression. And WTAP levels were positively correlated with levels of proliferation-related proteins, anti-apoptotic proteins, oncoproteins, and proteins important for stem cell function.
To gain more insight into the importance of WTAP, the investigators silenced its expression in K562 cells, HL-60 cells, OCI-AML3 cells, and primary AML cells.
“Knocking down this protein, WTAP, greatly suppressed proliferation and induced differentiation,” said study author Hima Bansal, PhD, of The University of Texas Health Science Center at San Antonio.
WTAP knockdown alone did not induce apoptosis, but it did enhance the apoptosis that occurred after the administration of etoposide.
The researchers also examined the role of WTAP in AML using mouse models. They found that tumors derived from WTAP-knockdown cells were significantly smaller and grew significantly slower than tumors derived from cells that expressed WTAP.
Finally, the investigators set out to determine why WTAP is overexpressed in AML. They noted that the Wilms’ tumor 1 (WT1) gene has an oncogenic role in leukemogenesis, and WTAP partners with WT1 to function as a switch gene, regulating the balance between cell quiescence and proliferation.
So the researchers decided to investigate Hsp90, a molecular chaperone that helps stabilize many oncoproteins, including WT1. And they found a direct interaction between Hsp90 and WTAP.
The Hsp90 inhibitor ganetespib promoted the degradation of WTAP in K562, MV4-11, and Kasumi-1 cell lines, as well as in leukemic blasts. In mice, ganetespib inhibited tumor growth.
And experiments suggested that ganetespib-mediated WTAP degradation is dependent on the ubiquitin-proteasome pathway. But the investigators said further research is needed to clarify WTAP’s mechanism of action.
Nevertheless, they believe the results of this research indicate that WTAP could be a promising therapeutic target for AML. ![]()

Credit: Rhoda Baer
The protein WTAP could play an important role in the development of acute myeloid leukemia (AML), according to new research.
Investigators discovered that AML cells have higher-than-normal levels of WTAP.
But silencing WTAP expression in leukemic cells can suppress proliferation and induce differentiation.
And, in mouse models of AML, knocking down WTAP can reduce tumor growth.
The researchers recounted these findings in a letter to Leukemia.
The team first uncovered high levels of WTAP in AML cells compared to normal peripheral blood mononuclear cells. And they found evidence to suggest that this contributes to abnormal cell behavior.
WTAP levels were not associated with individual cytogenetic abnormalities, but FLT3-ITD and NPM1 mutations were significantly correlated with WTAP expression. And WTAP levels were positively correlated with levels of proliferation-related proteins, anti-apoptotic proteins, oncoproteins, and proteins important for stem cell function.
To gain more insight into the importance of WTAP, the investigators silenced its expression in K562 cells, HL-60 cells, OCI-AML3 cells, and primary AML cells.
“Knocking down this protein, WTAP, greatly suppressed proliferation and induced differentiation,” said study author Hima Bansal, PhD, of The University of Texas Health Science Center at San Antonio.
WTAP knockdown alone did not induce apoptosis, but it did enhance the apoptosis that occurred after the administration of etoposide.
The researchers also examined the role of WTAP in AML using mouse models. They found that tumors derived from WTAP-knockdown cells were significantly smaller and grew significantly slower than tumors derived from cells that expressed WTAP.
Finally, the investigators set out to determine why WTAP is overexpressed in AML. They noted that the Wilms’ tumor 1 (WT1) gene has an oncogenic role in leukemogenesis, and WTAP partners with WT1 to function as a switch gene, regulating the balance between cell quiescence and proliferation.
So the researchers decided to investigate Hsp90, a molecular chaperone that helps stabilize many oncoproteins, including WT1. And they found a direct interaction between Hsp90 and WTAP.
The Hsp90 inhibitor ganetespib promoted the degradation of WTAP in K562, MV4-11, and Kasumi-1 cell lines, as well as in leukemic blasts. In mice, ganetespib inhibited tumor growth.
And experiments suggested that ganetespib-mediated WTAP degradation is dependent on the ubiquitin-proteasome pathway. But the investigators said further research is needed to clarify WTAP’s mechanism of action.
Nevertheless, they believe the results of this research indicate that WTAP could be a promising therapeutic target for AML. ![]()

Credit: Rhoda Baer
The protein WTAP could play an important role in the development of acute myeloid leukemia (AML), according to new research.
Investigators discovered that AML cells have higher-than-normal levels of WTAP.
But silencing WTAP expression in leukemic cells can suppress proliferation and induce differentiation.
And, in mouse models of AML, knocking down WTAP can reduce tumor growth.
The researchers recounted these findings in a letter to Leukemia.
The team first uncovered high levels of WTAP in AML cells compared to normal peripheral blood mononuclear cells. And they found evidence to suggest that this contributes to abnormal cell behavior.
WTAP levels were not associated with individual cytogenetic abnormalities, but FLT3-ITD and NPM1 mutations were significantly correlated with WTAP expression. And WTAP levels were positively correlated with levels of proliferation-related proteins, anti-apoptotic proteins, oncoproteins, and proteins important for stem cell function.
To gain more insight into the importance of WTAP, the investigators silenced its expression in K562 cells, HL-60 cells, OCI-AML3 cells, and primary AML cells.
“Knocking down this protein, WTAP, greatly suppressed proliferation and induced differentiation,” said study author Hima Bansal, PhD, of The University of Texas Health Science Center at San Antonio.
WTAP knockdown alone did not induce apoptosis, but it did enhance the apoptosis that occurred after the administration of etoposide.
The researchers also examined the role of WTAP in AML using mouse models. They found that tumors derived from WTAP-knockdown cells were significantly smaller and grew significantly slower than tumors derived from cells that expressed WTAP.
Finally, the investigators set out to determine why WTAP is overexpressed in AML. They noted that the Wilms’ tumor 1 (WT1) gene has an oncogenic role in leukemogenesis, and WTAP partners with WT1 to function as a switch gene, regulating the balance between cell quiescence and proliferation.
So the researchers decided to investigate Hsp90, a molecular chaperone that helps stabilize many oncoproteins, including WT1. And they found a direct interaction between Hsp90 and WTAP.
The Hsp90 inhibitor ganetespib promoted the degradation of WTAP in K562, MV4-11, and Kasumi-1 cell lines, as well as in leukemic blasts. In mice, ganetespib inhibited tumor growth.
And experiments suggested that ganetespib-mediated WTAP degradation is dependent on the ubiquitin-proteasome pathway. But the investigators said further research is needed to clarify WTAP’s mechanism of action.
Nevertheless, they believe the results of this research indicate that WTAP could be a promising therapeutic target for AML. ![]()
Mutant HSCs appear to drive AML
A new study has shown that hematopoietic stem cells (HSCs) can acquire mutations in DNMT3A, and this may be the first step in initiating acute myeloid leukemia (AML).
These HSCs also appear to be a means of treatment resistance and may trigger relapse in patients with AML, investigators reported in Nature.
“Our discovery lays the groundwork to detect and target the pre-leukemic stem cell and thereby potentially stop the disease at a very early stage, when it may be more amenable to treatment,” said study author John Dick, PhD, of the University of Toronto in Ontario, Canada.
“Now, we have a potential tool for earlier diagnosis that may allow early intervention before the development of full AML. We can also monitor remission and initiate therapy to target the pre-leukemic stem cell to prevent relapse.”
Dr Dick and his colleagues analyzed 71 samples from AML patients and discovered that 17 of them (24%) carried mutations in DNMT3A. Fifteen of those samples (88%) also had mutated NPM1.
Both mutations were present in patients’ blasts. But 12 patients (70.5%) had T cells that contained DNMT3A mutations but no NPM1 mutations. FLT3-ITD mutations were also present in blasts but not T cells in 2 patients.
These results suggest DNMT3A mutations arise earlier than NPM1 and FLT3-ITD mutations, the researchers said.
To determine the origin of mutated DNMT3A, they analyzed hematopoietic stem and progenitor cell populations from 11 patients with DNMT3A and NPM1 mutations.
While both types of mutations were present in CD33+ blasts, mutant DNMT3A was present without mutant NPM1 across the spectrum of mature and progenitor cell populations.
Experiments in mice revealed that DNMT3A-mutant HSCs had a multilineage repopulation advantage over non-mutant HSCs. This, the investigators said, establishes the mutant cells as pre-leukemic HSCs.
The team also found the pre-leukemic HSCs in samples taken from AML patients in remission, which showed that the cells survived chemotherapy.
The researchers therefore concluded that DNMT3A mutations arise early in AML evolution and lead to a clonally expanded pool of pre-leukemic HSCs from which AML develops.
“By peering into the ‘black box’ of how cancer develops during the months and years prior to when it is first diagnosed, we have demonstrated a unique finding,” Dr Dick said. “People tend to think relapse after remission means chemotherapy didn’t kill all the cancer cells.”
“Our study suggests that, in some cases, the chemotherapy does, in fact, eradicate AML. What it does not touch are the pre-leukemic stem cells that can trigger another round of AML development and, ultimately, disease relapse.”
Dr Dick believes this finding could spawn accelerated drug development to specifically target DNMT3A. The discovery should also provide impetus for researchers to look for pre-cancerous cells in AML patients with other mutations. ![]()
A new study has shown that hematopoietic stem cells (HSCs) can acquire mutations in DNMT3A, and this may be the first step in initiating acute myeloid leukemia (AML).
These HSCs also appear to be a means of treatment resistance and may trigger relapse in patients with AML, investigators reported in Nature.
“Our discovery lays the groundwork to detect and target the pre-leukemic stem cell and thereby potentially stop the disease at a very early stage, when it may be more amenable to treatment,” said study author John Dick, PhD, of the University of Toronto in Ontario, Canada.
“Now, we have a potential tool for earlier diagnosis that may allow early intervention before the development of full AML. We can also monitor remission and initiate therapy to target the pre-leukemic stem cell to prevent relapse.”
Dr Dick and his colleagues analyzed 71 samples from AML patients and discovered that 17 of them (24%) carried mutations in DNMT3A. Fifteen of those samples (88%) also had mutated NPM1.
Both mutations were present in patients’ blasts. But 12 patients (70.5%) had T cells that contained DNMT3A mutations but no NPM1 mutations. FLT3-ITD mutations were also present in blasts but not T cells in 2 patients.
These results suggest DNMT3A mutations arise earlier than NPM1 and FLT3-ITD mutations, the researchers said.
To determine the origin of mutated DNMT3A, they analyzed hematopoietic stem and progenitor cell populations from 11 patients with DNMT3A and NPM1 mutations.
While both types of mutations were present in CD33+ blasts, mutant DNMT3A was present without mutant NPM1 across the spectrum of mature and progenitor cell populations.
Experiments in mice revealed that DNMT3A-mutant HSCs had a multilineage repopulation advantage over non-mutant HSCs. This, the investigators said, establishes the mutant cells as pre-leukemic HSCs.
The team also found the pre-leukemic HSCs in samples taken from AML patients in remission, which showed that the cells survived chemotherapy.
The researchers therefore concluded that DNMT3A mutations arise early in AML evolution and lead to a clonally expanded pool of pre-leukemic HSCs from which AML develops.
“By peering into the ‘black box’ of how cancer develops during the months and years prior to when it is first diagnosed, we have demonstrated a unique finding,” Dr Dick said. “People tend to think relapse after remission means chemotherapy didn’t kill all the cancer cells.”
“Our study suggests that, in some cases, the chemotherapy does, in fact, eradicate AML. What it does not touch are the pre-leukemic stem cells that can trigger another round of AML development and, ultimately, disease relapse.”
Dr Dick believes this finding could spawn accelerated drug development to specifically target DNMT3A. The discovery should also provide impetus for researchers to look for pre-cancerous cells in AML patients with other mutations. ![]()
A new study has shown that hematopoietic stem cells (HSCs) can acquire mutations in DNMT3A, and this may be the first step in initiating acute myeloid leukemia (AML).
These HSCs also appear to be a means of treatment resistance and may trigger relapse in patients with AML, investigators reported in Nature.
“Our discovery lays the groundwork to detect and target the pre-leukemic stem cell and thereby potentially stop the disease at a very early stage, when it may be more amenable to treatment,” said study author John Dick, PhD, of the University of Toronto in Ontario, Canada.
“Now, we have a potential tool for earlier diagnosis that may allow early intervention before the development of full AML. We can also monitor remission and initiate therapy to target the pre-leukemic stem cell to prevent relapse.”
Dr Dick and his colleagues analyzed 71 samples from AML patients and discovered that 17 of them (24%) carried mutations in DNMT3A. Fifteen of those samples (88%) also had mutated NPM1.
Both mutations were present in patients’ blasts. But 12 patients (70.5%) had T cells that contained DNMT3A mutations but no NPM1 mutations. FLT3-ITD mutations were also present in blasts but not T cells in 2 patients.
These results suggest DNMT3A mutations arise earlier than NPM1 and FLT3-ITD mutations, the researchers said.
To determine the origin of mutated DNMT3A, they analyzed hematopoietic stem and progenitor cell populations from 11 patients with DNMT3A and NPM1 mutations.
While both types of mutations were present in CD33+ blasts, mutant DNMT3A was present without mutant NPM1 across the spectrum of mature and progenitor cell populations.
Experiments in mice revealed that DNMT3A-mutant HSCs had a multilineage repopulation advantage over non-mutant HSCs. This, the investigators said, establishes the mutant cells as pre-leukemic HSCs.
The team also found the pre-leukemic HSCs in samples taken from AML patients in remission, which showed that the cells survived chemotherapy.
The researchers therefore concluded that DNMT3A mutations arise early in AML evolution and lead to a clonally expanded pool of pre-leukemic HSCs from which AML develops.
“By peering into the ‘black box’ of how cancer develops during the months and years prior to when it is first diagnosed, we have demonstrated a unique finding,” Dr Dick said. “People tend to think relapse after remission means chemotherapy didn’t kill all the cancer cells.”
“Our study suggests that, in some cases, the chemotherapy does, in fact, eradicate AML. What it does not touch are the pre-leukemic stem cells that can trigger another round of AML development and, ultimately, disease relapse.”
Dr Dick believes this finding could spawn accelerated drug development to specifically target DNMT3A. The discovery should also provide impetus for researchers to look for pre-cancerous cells in AML patients with other mutations.
Normal enzyme aids mutated FLT3 to fuel AML

Results of preclinical research suggest the wild-type version of SYK pairs with mutated FLT3 to promote progression of acute myelogenous leukemia (AML).
And this molecular partnership promotes AML cells’ resistance to FLT3 inhibitors.
However, adding a SYK inhibitor to the mix can override this resistance. In an animal model of AML, treatment with a combination of FLT3 and SYK inhibitors was significantly more effective than either inhibitor alone.
These findings, published in Cancer Cell, raise hopes that treatment strategies focusing on both enzymes simultaneously could improve outcomes for patients with FLT3-ITD AML.
“Patients whose AML cells express FLT3-ITD are among the highest-risk group of patients with AML,” said study author Kimberly Stegmaier, MD, of the Dana-Farber Cancer Institute in Boston. “Their AML is particularly difficult to treat.”
In 2009, researchers in Dr Stegmaier’s lab discovered that SYK, a kinase that had attracted attention for its role in other malignancies, could be a potential drug target in AML. Unlike other cancer-associated kinases, SYK rarely undergoes mutations or other genomic alterations in cancer cells, remaining in its wild-type form.
With the current study, Dr Stegmaier and her colleagues set out to better understand SYK’s role in AML. The team screened AML cell lines to reveal the full scope of the enzyme’s molecular interactions. And they found evidence of strong interactions between wild-type SYK and mutated FLT3, particularly FLT3-ITD.
“We wanted to understand the cooperative oncologic effects by which SYK contributes to AML,” Dr Stegmaier said. “The concept of a normal enzyme aiding a mutant one has not yet been widely explored, and so we were both surprised and pleased to see FLT3-ITD come up as a high-priority hit in our screens.”
Through experiments in cell lines, primary patient samples, and animal models, the researchers found that SYK and FLT3-ITD’s interactions are a key ingredient in the progression of myeloproliferative neoplasms into AML. AML cells’ continued growth after turning malignant also relied on these interactions.
In addition, the team found that SYK’s hyperactivated form can promote resistance to the FLT3-targeting drug quizartinib. However, a combination of quizartinib and the SYK inhibitor PRT062607 overcame this resistance, significantly increasing survival and reducing signs of disease in a FLT3-ITD AML mouse model.
Highlighting their findings’ clinical relevance, the researchers found strong SYK activity in cells from FLT3-ITD AML patients. The cells were also highly sensitive to SYK inhibition.
“These data affirm that SYK is an important target in AML,” Dr Stegmaier said. “They also suggest that interactions between oncologic kinases and SYK or other wild-type enzymes may contribute to resistance of kinase inhibitors more broadly.”
Dr Stegmaier added that, over the course of this research, the team has developed a suite of tools that could prove useful for future clinical studies of treatments with SYK inhibitors or SYK inhibitors in combination with FLT3 inhibitors.
“We have not only identified SYK as a candidate treatment target in AML, but we have also identified a specific population of patients with AML more likely to respond to SYK inhibitors: patients with FLT3 mutations,” she said.
“Moreover, we have developed tools for identifying patients with high levels of SYK and FLT3 activation and can monitor these 2 targets while patients are receiving treatment. Predictive biomarkers of response are becoming increasingly important in the development of effective clinical trials of targeted therapies.”

Results of preclinical research suggest the wild-type version of SYK pairs with mutated FLT3 to promote progression of acute myelogenous leukemia (AML).
And this molecular partnership promotes AML cells’ resistance to FLT3 inhibitors.
However, adding a SYK inhibitor to the mix can override this resistance. In an animal model of AML, treatment with a combination of FLT3 and SYK inhibitors was significantly more effective than either inhibitor alone.
These findings, published in Cancer Cell, raise hopes that treatment strategies focusing on both enzymes simultaneously could improve outcomes for patients with FLT3-ITD AML.
“Patients whose AML cells express FLT3-ITD are among the highest-risk group of patients with AML,” said study author Kimberly Stegmaier, MD, of the Dana-Farber Cancer Institute in Boston. “Their AML is particularly difficult to treat.”
In 2009, researchers in Dr Stegmaier’s lab discovered that SYK, a kinase that had attracted attention for its role in other malignancies, could be a potential drug target in AML. Unlike other cancer-associated kinases, SYK rarely undergoes mutations or other genomic alterations in cancer cells, remaining in its wild-type form.
With the current study, Dr Stegmaier and her colleagues set out to better understand SYK’s role in AML. The team screened AML cell lines to reveal the full scope of the enzyme’s molecular interactions. And they found evidence of strong interactions between wild-type SYK and mutated FLT3, particularly FLT3-ITD.
“We wanted to understand the cooperative oncologic effects by which SYK contributes to AML,” Dr Stegmaier said. “The concept of a normal enzyme aiding a mutant one has not yet been widely explored, and so we were both surprised and pleased to see FLT3-ITD come up as a high-priority hit in our screens.”
Through experiments in cell lines, primary patient samples, and animal models, the researchers found that SYK and FLT3-ITD’s interactions are a key ingredient in the progression of myeloproliferative neoplasms into AML. AML cells’ continued growth after turning malignant also relied on these interactions.
In addition, the team found that SYK’s hyperactivated form can promote resistance to the FLT3-targeting drug quizartinib. However, a combination of quizartinib and the SYK inhibitor PRT062607 overcame this resistance, significantly increasing survival and reducing signs of disease in a FLT3-ITD AML mouse model.
Highlighting their findings’ clinical relevance, the researchers found strong SYK activity in cells from FLT3-ITD AML patients. The cells were also highly sensitive to SYK inhibition.
“These data affirm that SYK is an important target in AML,” Dr Stegmaier said. “They also suggest that interactions between oncologic kinases and SYK or other wild-type enzymes may contribute to resistance of kinase inhibitors more broadly.”
Dr Stegmaier added that, over the course of this research, the team has developed a suite of tools that could prove useful for future clinical studies of treatments with SYK inhibitors or SYK inhibitors in combination with FLT3 inhibitors.
“We have not only identified SYK as a candidate treatment target in AML, but we have also identified a specific population of patients with AML more likely to respond to SYK inhibitors: patients with FLT3 mutations,” she said.
“Moreover, we have developed tools for identifying patients with high levels of SYK and FLT3 activation and can monitor these 2 targets while patients are receiving treatment. Predictive biomarkers of response are becoming increasingly important in the development of effective clinical trials of targeted therapies.”

Results of preclinical research suggest the wild-type version of SYK pairs with mutated FLT3 to promote progression of acute myelogenous leukemia (AML).
And this molecular partnership promotes AML cells’ resistance to FLT3 inhibitors.
However, adding a SYK inhibitor to the mix can override this resistance. In an animal model of AML, treatment with a combination of FLT3 and SYK inhibitors was significantly more effective than either inhibitor alone.
These findings, published in Cancer Cell, raise hopes that treatment strategies focusing on both enzymes simultaneously could improve outcomes for patients with FLT3-ITD AML.
“Patients whose AML cells express FLT3-ITD are among the highest-risk group of patients with AML,” said study author Kimberly Stegmaier, MD, of the Dana-Farber Cancer Institute in Boston. “Their AML is particularly difficult to treat.”
In 2009, researchers in Dr Stegmaier’s lab discovered that SYK, a kinase that had attracted attention for its role in other malignancies, could be a potential drug target in AML. Unlike other cancer-associated kinases, SYK rarely undergoes mutations or other genomic alterations in cancer cells, remaining in its wild-type form.
With the current study, Dr Stegmaier and her colleagues set out to better understand SYK’s role in AML. The team screened AML cell lines to reveal the full scope of the enzyme’s molecular interactions. And they found evidence of strong interactions between wild-type SYK and mutated FLT3, particularly FLT3-ITD.
“We wanted to understand the cooperative oncologic effects by which SYK contributes to AML,” Dr Stegmaier said. “The concept of a normal enzyme aiding a mutant one has not yet been widely explored, and so we were both surprised and pleased to see FLT3-ITD come up as a high-priority hit in our screens.”
Through experiments in cell lines, primary patient samples, and animal models, the researchers found that SYK and FLT3-ITD’s interactions are a key ingredient in the progression of myeloproliferative neoplasms into AML. AML cells’ continued growth after turning malignant also relied on these interactions.
In addition, the team found that SYK’s hyperactivated form can promote resistance to the FLT3-targeting drug quizartinib. However, a combination of quizartinib and the SYK inhibitor PRT062607 overcame this resistance, significantly increasing survival and reducing signs of disease in a FLT3-ITD AML mouse model.
Highlighting their findings’ clinical relevance, the researchers found strong SYK activity in cells from FLT3-ITD AML patients. The cells were also highly sensitive to SYK inhibition.
“These data affirm that SYK is an important target in AML,” Dr Stegmaier said. “They also suggest that interactions between oncologic kinases and SYK or other wild-type enzymes may contribute to resistance of kinase inhibitors more broadly.”
Dr Stegmaier added that, over the course of this research, the team has developed a suite of tools that could prove useful for future clinical studies of treatments with SYK inhibitors or SYK inhibitors in combination with FLT3 inhibitors.
“We have not only identified SYK as a candidate treatment target in AML, but we have also identified a specific population of patients with AML more likely to respond to SYK inhibitors: patients with FLT3 mutations,” she said.
“Moreover, we have developed tools for identifying patients with high levels of SYK and FLT3 activation and can monitor these 2 targets while patients are receiving treatment. Predictive biomarkers of response are becoming increasingly important in the development of effective clinical trials of targeted therapies.”